THE COTTLE GROUP
Principal Investigator: Dr. Renee Cottle
RESEARCH
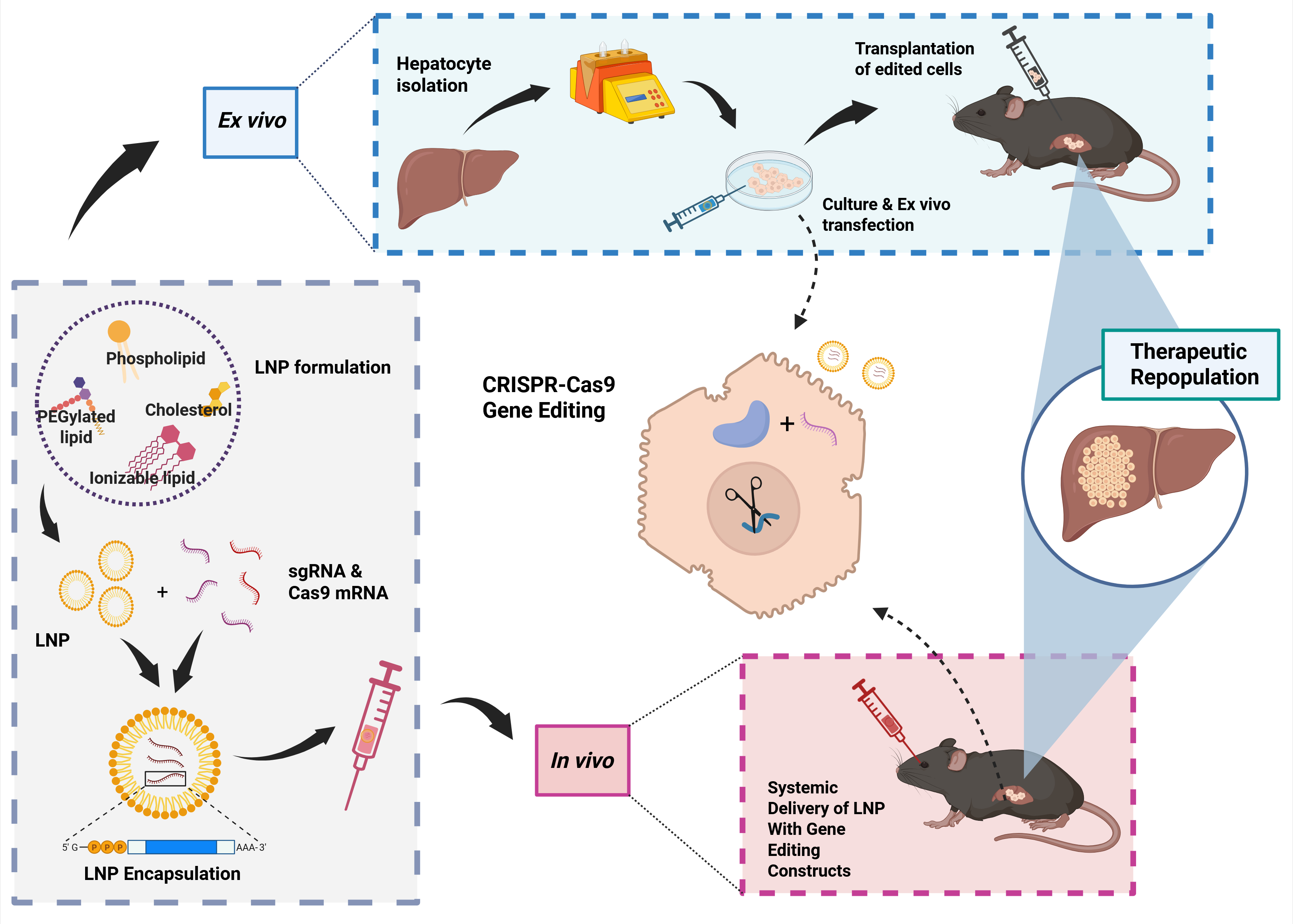
Nonviral gene delivery is increasingly recognized as a practical alternative to viral vectors, with growing evidence supporting its safety, scalability, and therapeutic potential. Among nonviral platforms, ionizable lipid nanoparticles (LNPs) have emerged as a leading technology for systemic in vivo gene delivery. However, their broader clinical application remains limited by dose-dependent toxicities, off-target biodistribution, immunogenicity, and challenges associated with repeat dosing. For liver-directed gene therapy, ex vivo gene editing of primary hepatocytes offers a complementary strategy that may improve therapeutic precision while minimizing systemic exposure and unintended off-target tissue targeting. Because liver targeting can be achieved through both in vivo and ex vivo approaches, these modalities can be deployed flexibly depending on the disease mechanism, therapeutic objectives, and translational considerations. Hereditary tyrosinemia type I (HTI) provides a well-established and experimentally tractable model for evaluating liver-directed gene editing strategies. The Cottle Lab has previously demonstrated that CRISPR-Cas9 delivered by electroporation can efficiently disrupt the gene encoding 4-hydroxyphenylpyruvate dioxygenase (HPD), a key enzyme in the tyrosine catabolism pathway, in hepatocytes, thereby creating a protective metabolic bypass that enables edited hepatocytes to survive, expand, and rescue the disease in a mouse model of HTI. We are investigating the use of ionizable LNPs for CRISPR-Cas9 delivery to primary hepatocytes, and comparing an in vivo systemic gene-editing strategy with an ex vivo gene-editing approach using integrated hepatocyte transplantation. We directly compare these delivery modalities to assess editing efficiency, hepatocyte engraftment, functional rescue, and key translational considerations, including gene-editing biodistribution, safety, immunogenicity, and long-term persistence of gene-edited hepatocytes.