THE COTTLE GROUP
Principal Investigator: Dr. Renee Cottle
RESEARCH
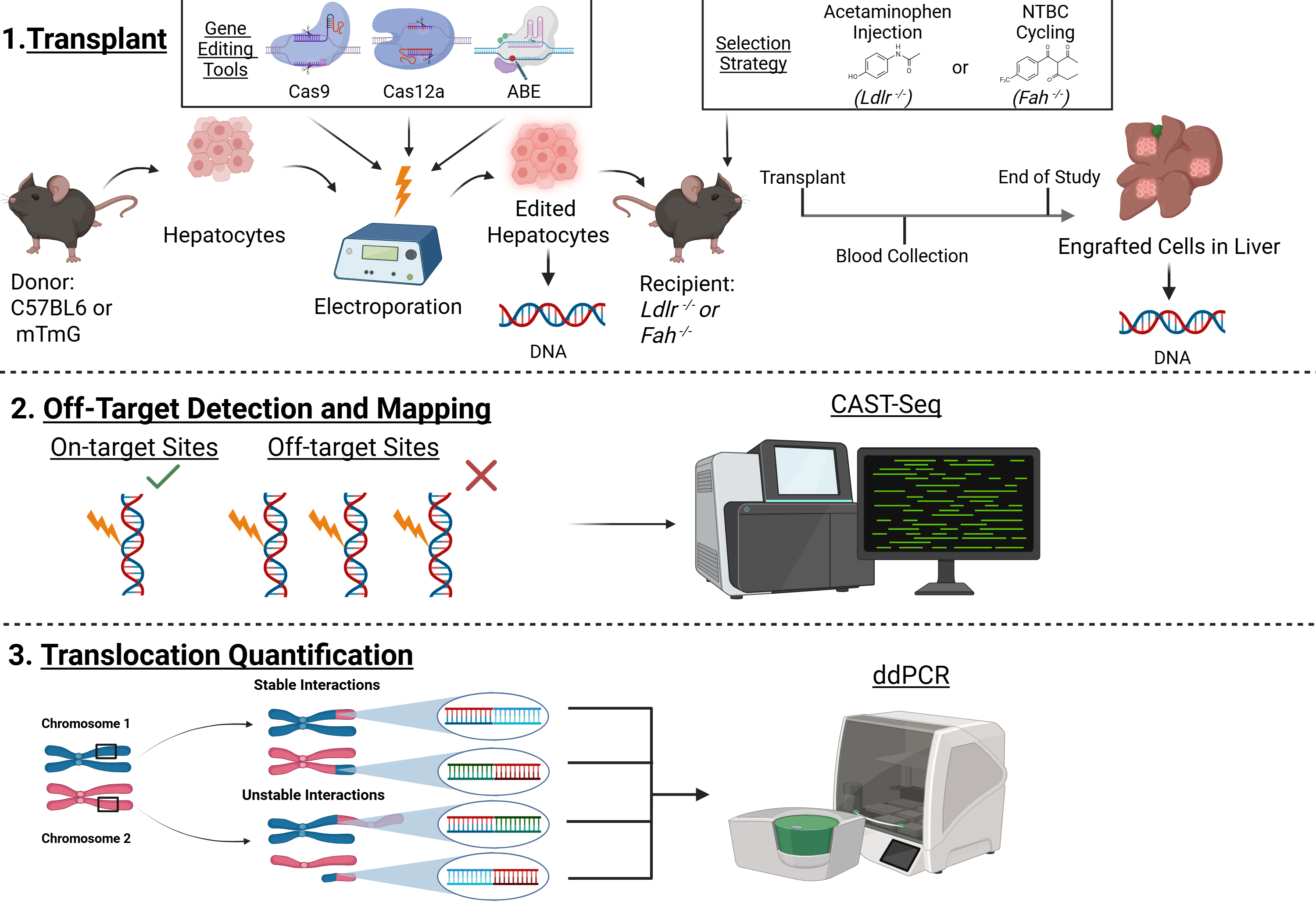
Because chromosomal translocations form primarily from double-strand DNA breaks (DSBs), alternative editing platforms that alter DNA without inducing DSBs may reduce genomic instability. While most clinical trials rely on CRISPR-Cas9 nucleases, other Cas effectors, such as Cas12a, and emerging CRISPR systems, like base editors, offer varied cutting kinetics or DSB-free nucleotide conversion, potentially reducing aberrant genomic rearrangements. However, no studies to date have systematically evaluated gene-editing specificity or the risk of chromosomal translocation events during multiplex CRISPR editing ex vivo. Critically, rigorous safety profiling of multiplex CRISPR systems in primary mouse and human hepatocytes, the target cell type for treatment of inherited metabolic liver diseases, remains largely unexplored. The Cottle Lab focuses on combining Cas nucleases systems for multiplex editing to achieve high on-target editing efficiency while reducing chromosomal translocations in both mouse and humanized mouse models. We are conducting studies to compare base editors and combinations of Cas nucleases in primary hepatocytes in culture and after engraftment and multiple cycles of replication in the liver in vivo. We are rigorously quantifying the frequencies of off-target editing and chromosomal translocations using droplet digital PCR (ddPCR) and CAST-seq, an unbiased genome-wide method that simultaneously detects off-target cleavage sites and translocation junctions. The aim of these studies is to establish a safety benchmark to guide the development of a clinically viable multiplex gene-editing strategy for inherited metabolic liver diseases.